Evolving Simple Organisms using a Genetic Algorithm and Deep Learning from Scratch with Python
TL;DR - Learn how to evolve a population of simple organisms each containing a unique neural network using a genetic algorithm.
Introduction
A few weeks ago I got pretty deep into a late night YouTube rabbit hole, and somewhere around evolving soft body robots, I came across this video (here). I’m not sure what if it was the peaceful background music or the hypnotizing motion of the dragonflies but I wanted to try and run the simulation on my local computer. After failing to find a GitHub repo for this, I decided to spend a few hours coding my own version in Python. I was pretty happy with the end results so I decided to turn it into a tutorial.
During this post, well code from scratch a simulation environment that contains organisms that must consume as much as food as possible in order to survive (click here to see the final results). If these organisms were people, they would be competitive eaters.

Neural Network and Navigation
Starting off with the basics, the organisms will be controlled by a simple, fully connected neural network. The input for the neural network will be a normalized value ranging from [-1, +1], representing the direction to the nearest food particle. This heading is calculated by taking the direction to the nearest food particle (which ranges from +/- 180 degrees) and dividing it by 180. Below are two navigation examples.
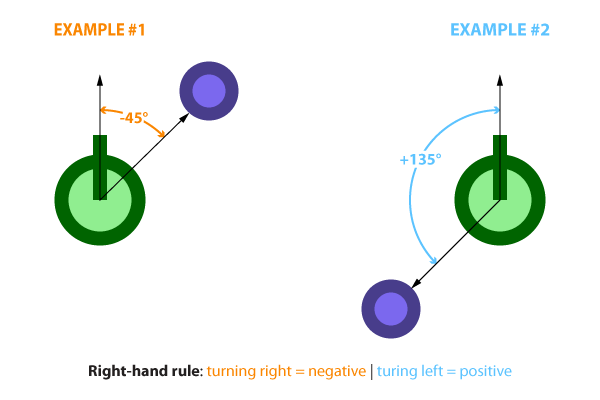
Because our single input ranges from [-1, +1] and our desired outputs also range from [-1, +1], the tanh will make the ideal activation function. Below is a diagram of the neural network with its inputs and outputs as well as its hidden layer. There exists many excellent tutorials on neural networks online so I wont go into great detail here. I recommend the following to those that are interested: here, here, here, and here.
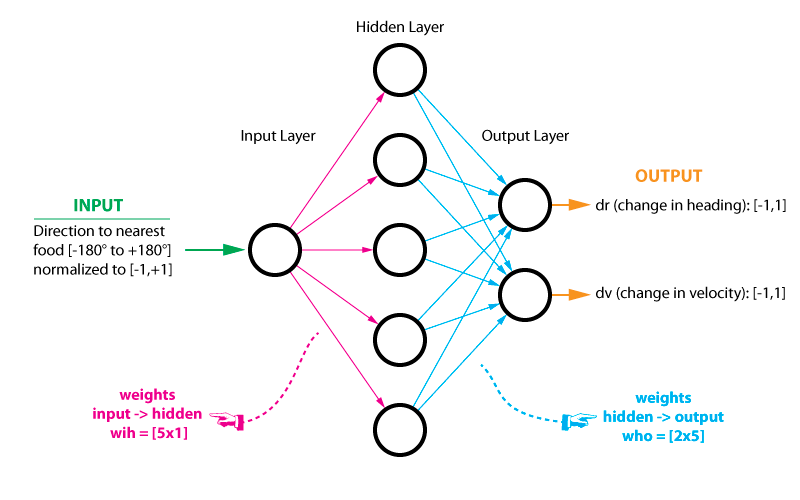
Since the neural network is nothing more than simple matrix multiplication, it only takes up a few lines of code thanks to NumPy. During the evolution process, it’s these weights that will be optimized. Lastly, I have set the default number of hidden nodes to five, but this can be increased or decreased to your liking.
Organism
The organism class is the star of the show here. It contains the neural network, as well as functions for updating its heading, velocity, and position. When an organism is initialized for the first time, its position, heading, velocity, acceleration, and neural network weights are all randomly generated.
class organism():
def __init__(self, settings, wih=None, who=None, name=None):
self.x = uniform(settings['x_min'], settings['x_max']) # position (x)
self.y = uniform(settings['y_min'], settings['y_max']) # position (y)
self.r = uniform(0,360) # orientation [0, 360]
self.v = uniform(0,settings['v_max']) # velocity [0, v_max]
self.dv = uniform(-settings['dv_max'], settings['dv_max']) # dv
self.d_food = 100 # distance to nearest food
self.r_food = 0 # orientation to nearest food
self.fitness = 0 # fitness (food count)
self.wih = wih
self.who = who
self.name = name
# NEURAL NETWORK
def think(self):
# SIMPLE MLP
af = lambda x: np.tanh(x) # activation function
h1 = af(np.dot(self.wih, self.r_food)) # hidden layer
out = af(np.dot(self.who, h1)) # output layer
# UPDATE dv AND dr WITH MLP RESPONSE
self.nn_dv = float(out[0]) # [-1, 1] (accelerate=1, deaccelerate=-1)
self.nn_dr = float(out[1]) # [-1, 1] (left=1, right=-1)
# UPDATE HEADING
def update_r(self, settings):
self.r += self.nn_dr * settings['dr_max'] * settings['dt']
self.r = self.r % 360
# UPDATE VELOCITY
def update_vel(self, settings):
self.v += self.nn_dv * settings['dv_max'] * settings['dt']
if self.v < 0: self.v = 0
if self.v > settings['v_max']: self.v = settings['v_max']
# UPDATE POSITION
def update_pos(self, settings):
dx = self.v * cos(radians(self.r)) * settings['dt']
dy = self.v * sin(radians(self.r)) * settings['dt']
self.x += dx
self.y += dy
Food
The only other class we need to create is the food class. This is a simple object that contains an x/y location and an energy value. This energy value will directly impact the organisms fitness. For now, this energy value remains constant and set at one, but it can be randomized or changed to anything if you feel like modifying it to. Additionally, a respawn function is present to regenerate the location of the food particle once it as been consumed by an organism. This keeps the total number of food particles within each simulation time step constant.
class food():
def __init__(self, settings):
self.x = uniform(settings['x_min'], settings['x_max'])
self.y = uniform(settings['y_min'], settings['y_max'])
self.energy = 1
def respawn(self,settings):
self.x = uniform(settings['x_min'], settings['x_max'])
self.y = uniform(settings['y_min'], settings['y_max'])
self.energy = 1
Evolution
The organism will be optimized using a genetic algorithm (GA) which falls under the larger umbrella of evolutionaty algorithms (EA). Genetic algorithms work by imitating the natural biological process of evolution by starting off with an initial population, and through selection, crossover, and mutation over many generations, an optimal solution emerges. With that said, EAs are not numerically guaranteed to find the global optimum but given proper initial conditions they are exceptionally good at searching through the design space and finding optimal or near optimal solutions. For this tutorial, the EA scheme will be as follows:
Selection - The simplest form of selection is called elitism, this is where the top n% of the current generations fittest individuals are selected and carried over to the next generation.
Crossover - Using the same pool of individuals selected during the elitism step previously, pairs of individuals will be randomly selected as parents and a new offspring will be generated. Because we're dealing with the weights of the neural network, the following equation will be used to crossover traits between the two parents to the resulting offspring:
$$ offspring=parent_1(a) + parent_2(1-a) $$ Where\( a \) is a randomly generated value between zero and one, and\( parent_1 \) , and\( parent_2 \) represent the weights of the neural network. Because there are two matrices that are being blending together (wih and who), the implementation will actually be as follows:$$ offspring\_wih=parent\_wih_1(a) + parent\_wih_2(1-a) $$ $$ offspring\_who=parent\_who_1(a) + parent\_who_2(1-a) $$ Mutation - Lastly, once the new offspring has been generated, a random number between zero and one will be generated. If this value is below the user initialized mutation threshold, mutation will occur. In the event that mutation does occur, a random weight from one of the two neural network weight matrices will be replaced with a random value that is within +/- 10% of the original value. This will create just a slight variation within the organism that could potentially lead to a fitter organism. The mutation effect is limited to +/- 10% of the original value because we want to avoid causing catastrophic failure within the neural network and potentially paralyzing the organism.
The diagram below shows just how simple this EA scheme is.
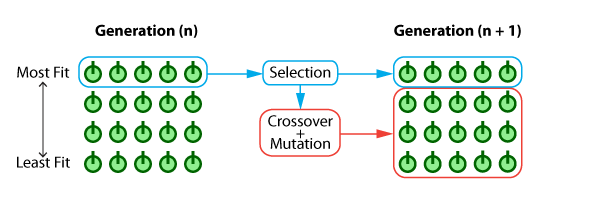
And finally, the codified EA routine can be seen below:
def evolve(settings, organisms_old, gen):
elitism_num = int(floor(settings['elitism'] * settings['pop_size']))
new_orgs = settings['pop_size'] - elitism_num
#--- GET STATS FROM CURRENT GENERATION ----------------+
stats = defaultdict(int)
for org in organisms_old:
if org.fitness > stats['BEST'] or stats['BEST'] == 0:
stats['BEST'] = org.fitness
if org.fitness < stats['WORST'] or stats['WORST'] == 0:
stats['WORST'] = org.fitness
stats['SUM'] += org.fitness
stats['COUNT'] += 1
stats['AVG'] = stats['SUM'] / stats['COUNT']
#--- ELITISM (KEEP BEST PERFORMING ORGANISMS) ---------+
orgs_sorted = sorted(organisms_old, key=operator.attrgetter('fitness'), reverse=True)
organisms_new = []
for i in range(0, elitism_num):
organisms_new.append(organism(settings, wih=orgs_sorted[i].wih, who=orgs_sorted[i].who, name=orgs_sorted[i].name))
#--- GENERATE NEW ORGANISMS ---------------------------+
for w in range(0, new_orgs):
# SELECTION (TRUNCATION SELECTION)
canidates = range(0, elitism_num)
random_index = sample(canidates, 2)
org_1 = orgs_sorted[random_index[0]]
org_2 = orgs_sorted[random_index[1]]
# CROSSOVER
crossover_weight = random()
wih_new = (crossover_weight * org_1.wih) + ((1 - crossover_weight) * org_2.wih)
who_new = (crossover_weight * org_1.who) + ((1 - crossover_weight) * org_2.who)
# MUTATION
mutate = random()
if mutate <= settings['mutate']:
# PICK WHICH WEIGHT MATRIX TO MUTATE
mat_pick = randint(0,1)
# MUTATE: WIH WEIGHTS
if mat_pick == 0:
index_row = randint(0,settings['hnodes']-1)
wih_new[index_row] = wih_new[index_row] * uniform(0.9, 1.1)
if wih_new[index_row] > 1: wih_new[index_row] = 1
if wih_new[index_row] < -1: wih_new[index_row] = -1
# MUTATE: WHO WEIGHTS
if mat_pick == 1:
index_row = randint(0,settings['onodes']-1)
index_col = randint(0,settings['hnodes']-1)
who_new[index_row][index_col] = who_new[index_row][index_col] * uniform(0.9, 1.1)
if who_new[index_row][index_col] > 1: who_new[index_row][index_col] = 1
if who_new[index_row][index_col] < -1: who_new[index_row][index_col] = -1
organisms_new.append(organism(settings, wih=wih_new, who=who_new, name='gen['+str(gen)+']-org['+str(w)+']'))
return organisms_new, stats
Simulation
The last critical piece of coding that’s needed is something to actually run the simulation. This simulate function will effectively represent the cost function for our optimization problem and will be called once every generation. The duration of each simulation is determined by dividing the total simulation time gen_time (in seconds) by the duration of the time step; dt. As an example, if the simulation length is set at 100 seconds, and dt is equal to 1/25th of a second, then there will be a total of 2500 timesteps that need to be simulated. During each time step, several key values will be updated:
Collision detection - Here we are checking for collisions between organisms and food particles. When a collision is detected, that organism will get its fitness function updated and the food particle will respawn at a new random location.
Determine closest food particle - This one is fairly self explanitory. For every organism, the closest food particle must be determined.
Determine direction to nearest food particle - Once the nearest food particle has been determined for each organism, the direction to that particle must be calculated.
Query neural network - Using the updated (and normalized) direction value, the neural network in each organism is quaried.
Update organism - Based on the respone from the neural network, the heading, velocity, and position are all updated.
Below is the simulation function:
def simulate(settings, organisms, foods, gen):
total_time_steps = int(settings['gen_time'] / settings['dt'])
#--- CYCLE THROUGH EACH TIME STEP ---------------------+
for t_step in range(0, total_time_steps, 1):
# PLOT SIMULATION FRAME
#if gen == settings['gens'] - 1 and settings['plot']==True:
if gen==49:
plot_frame(settings, organisms, foods, gen, t_step)
# UPDATE FITNESS FUNCTION
for food in foods:
for org in organisms:
food_org_dist = dist(org.x, org.y, food.x, food.y)
# UPDATE FITNESS FUNCTION
if food_org_dist <= 0.075:
org.fitness += food.energy
food.respawn(settings)
# RESET DISTANCE AND HEADING TO NEAREST FOOD SOURCE
org.d_food = 100
org.r_food = 0
# CALCULATE HEADING TO NEAREST FOOD SOURCE
for food in foods:
for org in organisms:
# CALCULATE DISTANCE TO SELECTED FOOD PARTICLE
food_org_dist = dist(org.x, org.y, food.x, food.y)
# DETERMINE IF THIS IS THE CLOSEST FOOD PARTICLE
if food_org_dist < org.d_food:
org.d_food = food_org_dist
org.r_food = calc_heading(org, food)
# GET ORGANISM RESPONSE
for org in organisms:
org.think()
# UPDATE ORGANISMS POSITION AND VELOCITY
for org in organisms:
org.update_r(settings)
org.update_vel(settings)
org.update_pos(settings)
return organisms
Putting it All Together
With all the major components having been created the final code can now be assembled. There are a few smaller functions I neglected to include in the above writeup because they’re so simple reading about them would take longer than just reading the code directly. I ended up using matplotlib for displaying the simulation. This is far from the most efficient or elligant solution so by default the visualization is turned off. Lastly, pasting the entire code would be a bit redundent at this point so download/fork it from the GitHub repo (here).
Results
When you run the entire code, the output should be something similar to this:
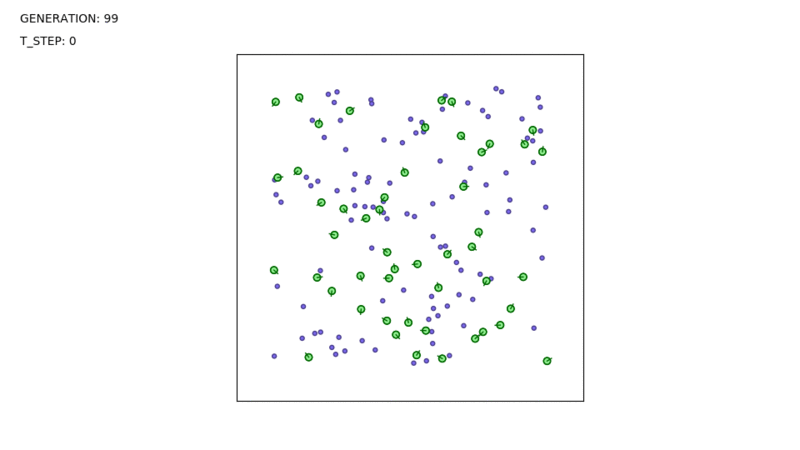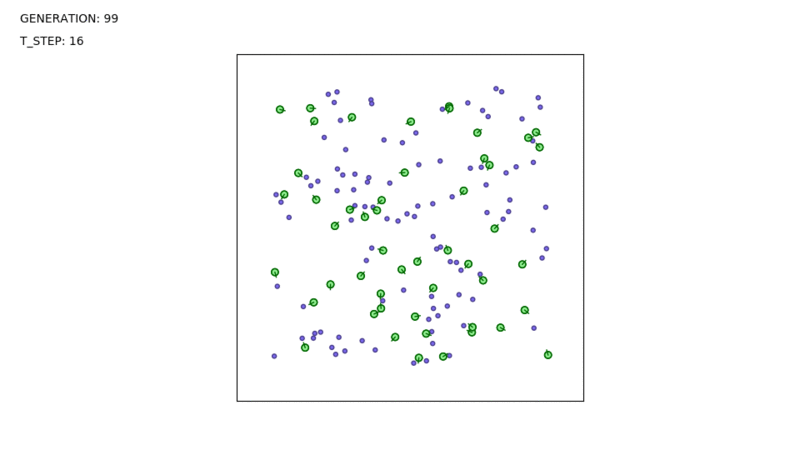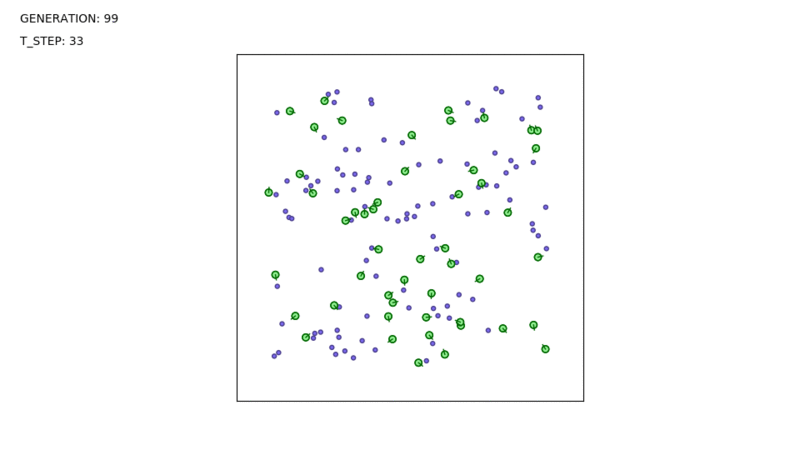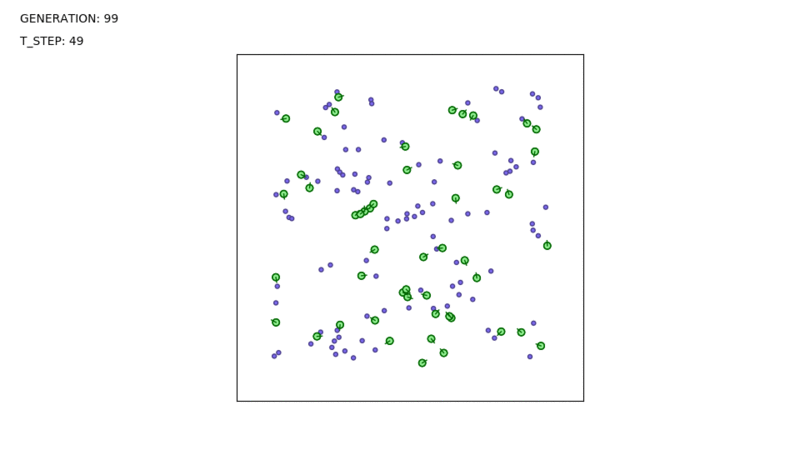
I hope this was helpful. If anything seems unclear, please let me know. Thanks for reading!